Bioinformatics consulting and analysis services
Based in Europe, Vidya BioSeq supports academic, clinical and biotech teams with the outsourcing of their omics data analyses.
Single-Cell RNA-Seq Analysis
scRNA-seq can now be applied to the vast majority of tissues and experimental contexts. Whether you work in oncology, immunology, virology or neuroscience, this technology opens up biological questions that bulk RNA-seq simply cannot address. We design workflows tailored to your objectives — to uncover rare cell subpopulations, identify novel molecular mechanisms, or characterise clinically or commercially relevant biomarkers.
- Rigorous, data-driven filtering — we do not apply arbitrary thresholds. Every filtering decision is justified by the actual distribution of your data, to avoid discarding biologically relevant cells.
- Validated cell type annotation — combining reference databases (CellMarker, PanglaoDB) with manual biological curation. Every cluster is documented and traceable for peer review.
- Cell dynamics — developmental trajectory reconstruction, pseudotime inference and RNA velocity to model the state transitions hidden in your data.
- From differential expression to biology — we don't deliver a gene list. We place results in their functional context (GO, KEGG, Reactome) and translate them into interpretable biological signatures.
Spatial Transcriptomics
scRNA-seq tells you which cells are present. Spatial transcriptomics tells you where they are — and how their immediate microenvironment shapes their transcriptional state.
We work with all major platforms — Visium (10x Genomics), Xenium, MERFISH, Slide-seq, CosMx — and adapt the analytical pipeline to the resolution and nature of your tissue samples.
- Tissue segmentation and spatial clustering — identification of transcriptomic domains and anatomical structures without prior manual annotation.
- Cell type deconvolution — integration with your scRNA-seq data to resolve the cellular composition of each spot or spatially resolved cell.
- Spatially resolved differential expression — comparison between regions of interest defined histologically or algorithmically, accounting for the spatial dependency of observations.
- Cellular niche analysis — identification of recurrent co-localisation patterns and spatially anchored cell-cell interactions.
- Multimodal integration — alignment of spatial data with scRNA-seq, histological imaging or proteomic data for a comprehensive, multi-layered interpretation.
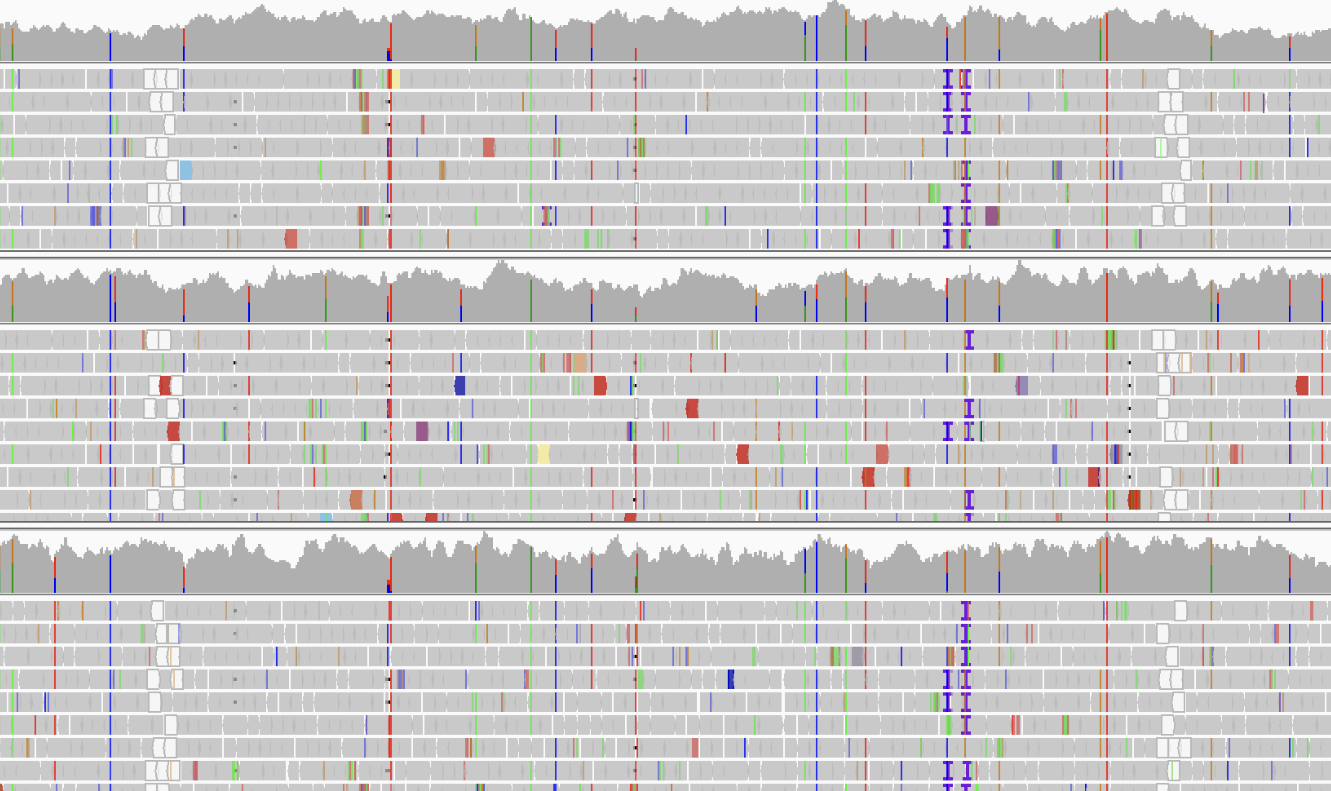
Genomics and Variant Analysis
Identifying a variant is only the first step. Determining whether it is pathogenic and functionally relevant is the core of the biological or clinical question. Our DNA-seq pipelines are built to reach that interpretive step — not just produce a raw VCF file.
- Constitutional and somatic variant detection — SNPs, indels, CNVs and structural variants (SVs), with tools and parameters adapted to each context, germline or somatic.
- Functional annotation and prioritisation — each variant is cross-referenced against major databases (ClinVar, gnomAD, OMIM) and prioritised according to its functional relevance and population frequency.
- ACMG classification — variant interpretation following American College of Medical Genetics criteria, for results that are directly usable in a diagnostic context.
- Trio and population analyses — de novo variant identification, familial segregation studies and allele frequency analyses in cohorts.
- Multi-omic integration — option to integrate genomic data with clinical RNA-seq for enhanced variant interpretation.
Epigenetics
Epigenomics addresses a question that genomics alone cannot answer: why do two cells with identical genomes behave differently? Chromatin accessibility, transcription factor occupancy, DNA methylation — these regulatory mechanisms are often at the heart of the pathological or developmental phenotypes you are trying to understand.
- ATAC-seq — open chromatin region detection, transcription factor footprinting and differential accessibility analysis between conditions or cell types.
- ChIP-seq — chromatin-binding protein occupancy mapping, signal quantification, motif analysis and multi-sample comparisons.
- DNA methylation — bisulfite-seq analysis, genomic annotation of differentially methylated regions, integration with gene expression data.
- Multi-omic integration — combining epigenomic data with RNA-seq to directly link active regulatory regions to their transcriptional impact.

Experimental Design Support
Reliable data starts with a well-designed experiment.
A bioinformatics consulting firm does more than analyse existing data. We work with research teams to optimise the bioinformatic and statistical aspects of their experimental protocol before data acquisition — ensuring scientific rigour, analytical interpretability and cost efficiency from the outset.
Our services include:
- Power analysis and sample size estimation for omics studies
- Randomisation strategies and batch effect mitigation
- Selection of appropriate controls, replicates and time points
- Definition of analysis-ready metadata structures (FAIR principles)

Scientific Writing Support
We help researchers turn complex analyses into clear, publication-ready results.
Our services include:
- Drafting and structuring methods and results sections
- Production of high-quality figures and tables
- Critical review of bioinformatic interpretations
- Writing responses to reviewers
- Preparation of supplementary materials (code, detailed methods, data summaries)
We act as genuine scientific collaborators, not merely service providers — to ensure your work is thoroughly documented, reproducible and impactful.

Custom Pipeline Development
Beyond standardised workflows, we offer:
- Bespoke pipeline development (Snakemake, Nextflow, WDL)
- Statistical modelling, machine learning and dimensionality reduction
- Database design and implementation for biological data management
- Interactive dashboards and reproducible reports (R Markdown, Shiny, Quarto, Jupyter Notebook)
- API development for integration with existing research infrastructure
- User-friendly interfaces designed for non-technical researchers
We work closely with experimental teams to align biological objectives with analytical feasibility — reducing back-and-forth, improving reproducibility and maximising the value of your data from day one.

Script Review and Optimisation
Have you inherited scripts from colleagues, former PhD students or postdocs and want to use them as a starting point for new work? Or bring them up to the standard required for publication?
We can help with all major languages used in bioinformatics, including:
- High-level scripting languages: Perl, Julia, R and Python
- Compiled low-level languages: C, C++ and Rust
- Workflow languages: Snakemake and Nextflow
- Other languages: SQL, HTML, CSS…

Frequently Asked Questions
What are the advantages of working with an external bioinformatics provider?
Engaging a specialist firm gives you immediate access to deep expertise without the delays and costs of recruitment — training, salary, software licences, infrastructure. You also benefit from an independent perspective on your data, which is often more rigorous than in-house analysis where confirmation bias can influence interpretation. Working with us means your team stays focused on the science while we handle the analytical complexity.
Is outsourcing suitable for exploratory analyses?
Yes — and it is often where external support adds the most value. A well-conducted exploratory analysis prevents teams from committing to a costly direction based on artefacts or methodological bias. We are regularly brought in during exploratory phases to help teams understand what their data actually contains before formalising hypotheses. We adapt the scope of analysis and deliverables accordingly — less formal, more iterative.
Outsourcing vs in-house bioinformatics — how do you choose?
The two approaches are not mutually exclusive. In-house capacity makes sense when your team generates data continuously and runs standardised, repetitive analyses. Outsourcing becomes the right choice for complex or one-off projects requiring highly specialised expertise — scRNA-seq, spatial transcriptomics, multi-omic integration — that you don't use frequently enough to justify a dedicated hire. Many of our clients combine both: an internal bioinformatics team for routine work, and Vidya BioSeq for high-value analyses.
How is the confidentiality of my research protected?
Confidentiality is a non-negotiable requirement before any collaboration begins. We sign a non-disclosure agreement (NDA) prior to any exchange of data or unpublished results. Data is transferred through secure channels and stored on access-restricted servers. We never publish, share or reuse client data without explicit written consent. Where your institution has specific security protocols — particularly in clinical or hospital settings — we comply with them fully.
What information should I prepare before getting in touch?
To help us understand your needs, it is useful to have a clear picture of your research objectives, the type of data you have or plan to generate, the analyses you are considering, your timeline, and any specific challenges you are facing. This allows us to provide a more accurate assessment and a tailored proposal.
What are typical project turnaround times?
Timelines vary with the complexity of the analyses. Smaller projects can be completed within one to two weeks; more complex analyses may take several months. We provide a detailed timeline estimate during the initial consultation and can adjust our workflow to meet urgent deadlines where possible.
Do you offer training for our team?
Yes. We offer hands-on workshops and knowledge transfer sessions as part of our service offering. These can be tailored to your team's existing skill level and areas of interest.
Looking for a tailored solution?
Our team can develop bioinformatics approaches designed around your specific research challenges.
Confidential discussion — response within 48 hours.
Get in touch